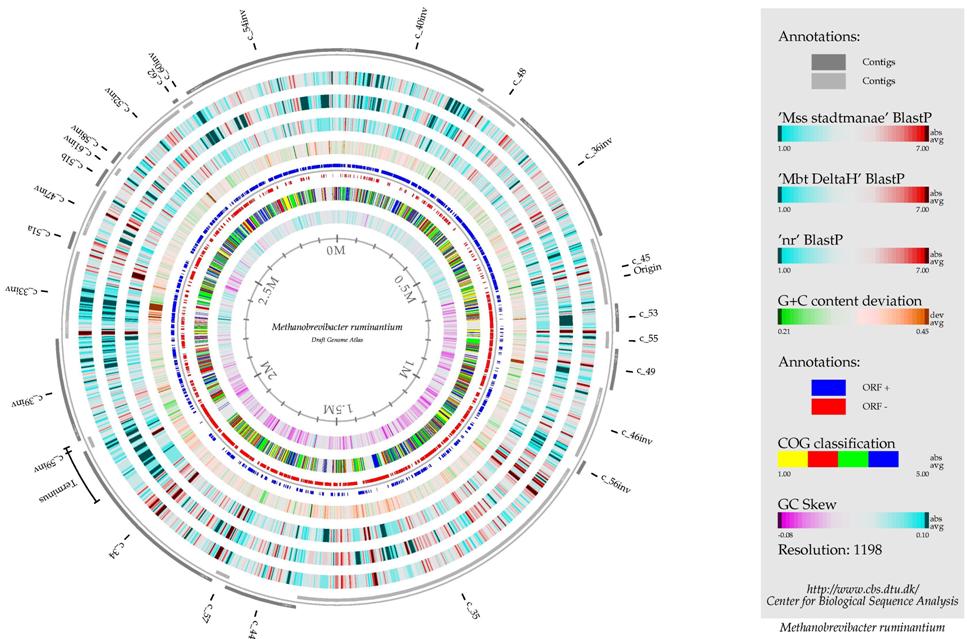
Today, the first map of a rumen methanogen DNA sequence was published in the journal PLoS One, giving scientists worldwide a major new opportunity to identify methods for cutting methane emissions from cows.
There was much excitement in late 2008 when it was announced that a team of PGgRC/AgResearch scientists were the first in the world to map the DNA sequence of a rumen methanogen.
Rumen methanogens are the bacteria responsible for the methane produced by livestock. The bacteria – of which there are a number of species – live in the gut of ruminant livestock, removing the hydrogen and carbon dioxide released as grass and other plant materials are broken down. The byproduct of this process, however, is large amounts of methane: one of the most potent greenhouse gases known.
The objective behind sequencing a rumen methanogen – in this case Methanobrevibacter ruminantium, a bacterium with 2200 genes and almost 3 million basepairs – is to figure out how to selectively knock them out in ways which will not damage other, beneficial bacteria. Possible approaches are vaccines, drenches or even changing forage.
The team, headed by Agresearch’s Dr Graeme Attwood, has been working on the project for over five years, but the PGgRC estimates it will still be a few years until any practical means of reducing methane can be developed.
The SMC asked local scientists to comment on the significance of the sequence and its release.
Dr Graeme Attwood, AgResearch scientist and leader of the team responsible, comments:
“The publication of the Methanobrevibacter ruminantium genome is a significant step towards understanding methane formation in ruminant animals. Methane is an important greenhouse gas (GHG), having a global warming potential 21 times that of carbon dioxide and makes up 66% of GHG emissions from agricultural activities in New Zealand.
“Enteric fermentation in the forestomach of ruminant livestock is the main source of methane, where it is produced by the action of a group of microorganisms called methanogens. Methanobrevibacter ruminantium is a prominent rumen methanogen and its 2.93 Mb (million basepairs) genome sequence has given us a detailed insight into its lifestyle in the rumen and allows us to identify genes and proteins as targets for methane reduction strategies.
“The Rumen Microbiology group based at Grasslands and the Vaccine Development group at the Hopkirk Research Institute are now using the gene sequence information in two different ways; an immune system approach which identifies parts of methanogens that aims to stimulate salivary antibodies to work against these microbes in the rumen; and a chemogenomics approach which uses small molecule inhibitors that target essential methanogen enzymes.
“The methanogen genome sequence also revealed several unexpected features; a complete prophage sequence encoding what appears to be a function bacteriophage (virus) and two non-ribosomal peptide synthetases which are the first reported in an archaeal species. We are excited by the possibilities that this genome sequence has unveiled and we are using the sequence information to work towards finding methane mitigation technologies for ruminant animals.”
Mark Aspin, Manager of the Pastoral Greenhouse Gas Research Consortium (PGgRC), comments:
“The PGgRc invested in this approach to develop the fundamental understanding of the Archaea methanogen- the organism for transforming hydrogen produced during digestion in the rumen to Methane.This process is critical to ensuring the rumen fermentation is highly efficient.
“The final publication is the result of a great partnership between the Farming Industry and Science over many years. The genome sequence gives us a blueprint of these unique organisms and will, we believe, enable specific targeting of methanogens which are distinctly different from the microbes responsible for fibre degradation (and hence digestion for the ruminant animal) so that we can reduce the emissions without adversely affecting their productivity.
“Apart from being able to specifically target the organism, this knowledge also improves researcher’s ability to understand and monitor rumen function and hence lead to improvements in that function and (hopefully) increased productivity.
“We will use this understanding across all of our research programmes where we are making changes to the rumen environment such as breeding or diet changes, so this investment will continue to deliver benefit to our research.
“This is only one of many species of methanogens in the rumen, the consortium has commenced further sequencing of other predominant methanogens so that we are confident that targets identified in the initial genome have wide application across the whole population of methanogens.
“We are also cooperating with other international groups that are sequencing rumen methanogens. Publication of the sequence will enable all researchers across the world to better understand these organisms and develop solutions to reduce methane emissions from ruminant livestock.”
Professor Jacqueline Rowarth, Director of Agriculture at Massey University, comments:
“The sequencing is a major breakthrough, not just for the scientists involved, but for New Zealand and all countries concerned about livestock contribution to GHG. You have to know what you’ve got before you can do anything about it. Now NZ knows the gene sequence, it can target specific actions for mitigation of methane production.
“NO other country is quite so concerned about the impact of cows eructating (burping), and so it is appropriate that this work has been done here. The breakthrough supports the belief that NZ is leading the world in this area and so is an appropriate country to lead the Global Alliance announced at Copenhagen.”
Professor Hugh Morgan, of the Department of Biological Sciences at the University of Waikato, comments:
“The paper represents a significant advance in efforts to understand and control methane emissions by ruminants. It provides no direct answers to these problems in itself, but represents a foundation stone on which hypotheses can be constructed and tested. It already provides a framework on which two competing approaches for methane control can be tested i.e. an immunogenetic approach whereby the host animal will be induced produce to broad spectrum antibodies to limit or inhibit methanogen growth, or the production and administration of specific inhibitors of methanogen metabolism.
“The complexity and diversity of the rumen microbial flora has engendered a feeling that it is an imperturbable steady state system; alter a feature or eliminate a species of bacterium and there would be others waiting to occupy the niche and carry out the same essential function. The manuscript reveals some intriguing features of the methanogen for adaptation to a rumen environment which suggest this not to be the case.
“The improved growth of this methanogen on short chain alcohols (though not supporting growth themselves) indicates a unique adaptation to this environment which can open potential control options, and if this growth stimulation is unique to ruminant methanogens, then the likelihood that other methanogen species occupying this niche is diminished. Similarly, the role of surface adhesin proteins whose putative function is to associate the methanogen with its hydrogen producing symbiont also points to a potential avenue of control. The authors identify likely proteins to target as vaccine agents, which is possibly the most promising method of control for implementation in the near future.
“The major challenge would seem to be to ensure that any vaccine is sufficiently broad-based to affect all rumen methanogens and not just the dominant species – this can now be tested directly and when other rumen methanogens are sequenced then better designed vaccines can be produced.
“As with all genome sequences there are surprises whose significance is difficult to forsee. In this instance, the role of non-ribosomally produced peptides and some expressed viral proteins are intriguing, but might yet have key roles to play in our understanding of rumen metabolism.”
Further Information
To talk to any of the experts quoted above contact the Science Media Centre on tel: 04 499 5476 or email: smc@sciencemediacentre.co.nz.
Notes to Editors
The Science Media Centre (SMC) is an independent source of expert comment and information for journalists covering science and technology in New Zealand. Our aim is to promote accurate, bias-free reporting on science and technology by helping the media work more closely with the scientific community. The SMC is an independent centre established by the Royal Society of New Zealand with funding from the Ministry of Research, Science and Technology. The views expressed in this Science Alert are those of the individuals and organisations indicated and do not reflect the views of the SMC or its employees. For further information about the centre, or to offer feedback, please email us at smc@sciencemediacentre.co.nz.